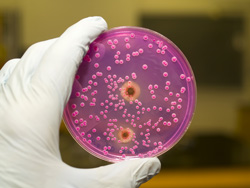
Budding yeast to reveal the secrets of DNA repair
Meiosis is a special kind of cell division that helps sexually reproducing populations adapt and evolve. Meiotic recombination begins when programmed DNA double-strand breaks (DSBs) are set in action by the Spo11 protein. This is an enzyme that cuts the DNA molecule due to receive new genetic information. In the yeast Saccharomyces cerevisiae, it has been found that the DSB sites don't just randomly place themselves along chromosomes; certain genomic areas are more likely to form DSBs than others. Histones also play a part in this process. These are the main protein components of chromatin found in cell nuclei of eukaryotes and are responsible for arranging DNA into its most basic structural form - the nucleosome. The EU-funded project 'Establishing the meiotic recombination-initiation epigenetic code in the yeast Saccharomyces cerevisiae' (EMRES) aimed to discover how histone modifications are involved in choosing and activating sites where meiotic recombination is set off. All work was done using the model organism S. cerevisiae (budding yeast). Researchers constructed a number of deletion mutants in order to block genes coding for histone-modifying enzymes. Meiosis was followed and extensively described from the meiotic S phase, DSB and crossover formation, sporulation efficiency through to spore viability. The 'chip-on-chip' technique, used to measure binding sites for proteins, was used for genome-wide distribution of H3K56ac. This marker is vital for the proper assembly of nucleosomes and maintaining genome stability during DNA replication. This mapping showed a random distribution process of the H3K56ac histone mark, and no correlation was found between the sites of meiotic DSB formation and H3K56ac. Having changed the original experimental strategy due to early study results, project partners developed a new, more adaptable means of examining histone modification during meiosis - a plasmid shuffling-based screen. After testing the screen, EMRES concluded that mutations of histone H3K4R and H3K4Q markers reiterate the effect of set1 deletion, which increases the capabilities of mutants following DNA damage. Following this, the research team was able to proceed with testing all the established histone mutants on the process of meiotic recombination.