Engineered bacteria hold out promise as vaccination ‘chassis’
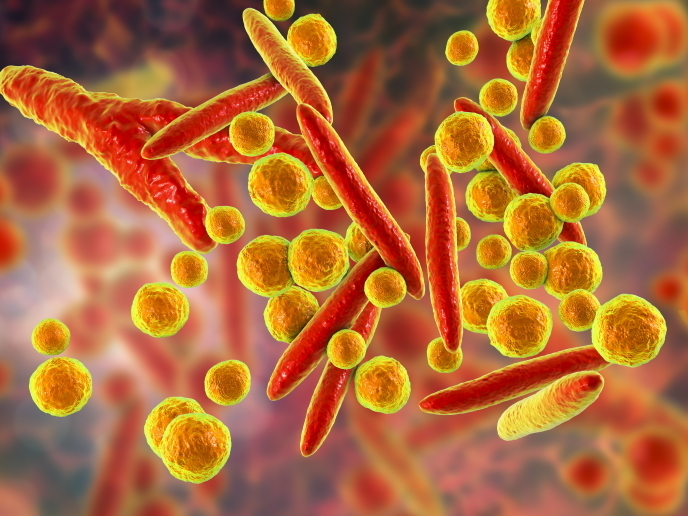
Bacteria are attractive therapeutic agents for vaccines. They can be engineered to carry the telltale molecular signatures (antigens) of different pathogens or cancer cells on their surface, alongside peptide adjuvants that boost the immune system response. Yet further development has been hampered by significant challenges. One problem is ensuring the immune system is stimulated by the added peptides and not endogenous proteins on the surface of the bacteria, which would compromise an effective immune response. The EU-supported MYCOCHASSIS project explored the use of single-celled mycoplasma pneumoniae(opens in new window) bacteria for vaccine development. The bacteria are normally pathogenic, causing respiratory disease in humans and livestock, but can be attenuated. Notably, they lack a cell wall. “Not having a cell wall makes it very easy to expose antigenic peptides and adjuvants on the bacterium’s surface – useful for vaccines. It also prevents the immune system from focusing on the cell wall, which contains lipopolysaccharides(opens in new window) that trigger inflammation,” explains project coordinator Luis Serrano from the Centre for Genomic Regulation(opens in new window), the project host. The project’s commercial partner MSD(opens in new window) is now testing and validating the engineered vaccines in animals, with a patent application forthcoming. The project has also resulted in the creation of a new company, Pulmobiotics(opens in new window), dedicated to using mycoplasma pneumoniae to treat human lung diseases.
Precision engineering
As researchers already knew a lot about the biology and pathogenicity of mycoplasma pneumoniae, Serrano’s team saw the potential of engineering these bacteria for disease treatment. Because the bacteria’s lack of a cell wall makes them easy to manipulate without compromising the immune response, “we thought vaccination would be one of the best medical interventions to try first,” adds Serrano. The team developed a better understanding of mycoplasma pneumoniae using the virtual model of the entire life cycle of the bacterium(opens in new window) developed by Markus Covert. This simulated interaction of DNA and cell components allowed the team to run scenarios to predict the likely immunological outcomes of molecular tweaks to the bacterium, ensuring efficacy and safety. “A model of a living system does not necessarily tell you that what you are doing is truly correct, but it can alert you to possible pitfalls in your design,” says Serrano. Based on what they learned from the modelling, the team started experimenting with transposons(opens in new window) – mobile elements of DNA that insert randomly in the genome. As these enabled the team to manipulate any DNA fragment of the bacteria’s genome, they were able to delete non-essential virulent genetic elements to create a non-pathogenic strain. Finally, a proof of concept was developed for animal vaccination. The best antigenic peptides of pathogenic bacteria which infect pig and cows were first determined, then expressed on the surface of mycoplasma pneumoniae as the vaccine chassis. “We observed a strong immune response against the exposed antigens in animals injected with the bacteria,” notes Serrano. As well as offering a non-pathogenic bacterium as a vaccination chassis, MYCOCHASSIS’s bacterium engineering approach could also be used to treat infectious human lung diseases such as COVID-19 and further in the future, lung cancer. Towards this end, the team are currently engineering mycoplasma pneumoniae to express immunomodulatory molecules(opens in new window), which typically confer immunity, as well as antibodies against different lung diseases.