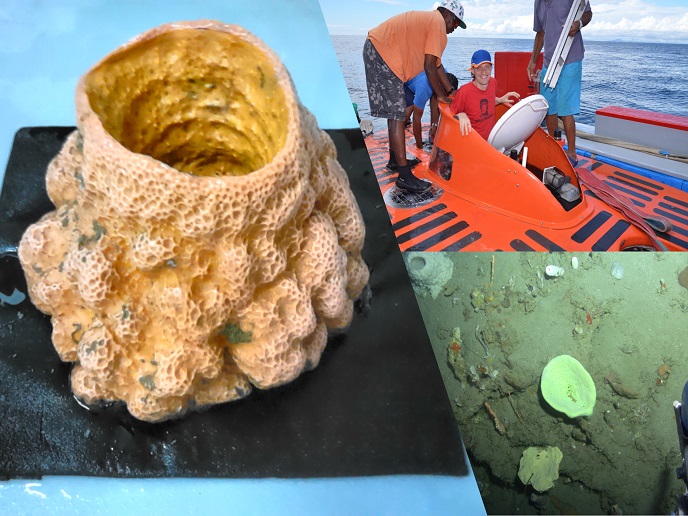
Shining light on ‘twilight zone’ sponges and their microbial cohabitants
Living organisms are a rich source of natural products that can benefit humans. Pharmaceuticals, nutraceuticals and skin care products are among the many commodities with increasingly natural and enhanced functionality thanks to biomolecules. With the support of the Marie Skłodowska-Curie Actions (MSCA) programme and high-tech genomics approaches, the COSMos project has provided an unprecedented data set pointing to the natural product potential of rarely studied mesophotic sponges and the microbes living in them.
A deep dive into the uncharted mesophotic zone
According to Michelle Schorn, MSCA Individual Fellow in the lab of Detmer Sipkema at Wageningen University(opens in new window), “sponges are renowned for their incredible chemical diversity and rich microbiomes (microbes that live in the sponges), which consistently provide the most novel natural products of all the marine invertebrates.” Sponges are primarily collected by scuba divers in shallow waters. The mesophotic zone, literally the ‘mid-light zone’, 30 to 150 m under the water’s surface, is difficult to access and largely unexplored. A serendipitous offer by Adriaan Schrier of Substation Curaçao(opens in new window) to join a sampling expedition to survey the mesophotic zone resulted in approximately 60 precious mesophotic sponge samples.
Metagenome-assembled genomes and biosynthetic gene clusters
Schorn got to work harnessing metagenome sequencing(opens in new window), a culture-independent genomic analysis method that enables the analysis of the combined genomes of organisms co-existing in a community by constructing metagenome-assembled genomes (MAGs). She used metagenomic sequencing to focus her ‘lens’ on biosynthetic gene clusters – physically clustered groups of genes that encode enzymes, which work together to make molecules or chemical structures – that are responsible for most natural products. “Less than 1 % of our biosynthetic gene clusters corresponded to those known, highlighting the novelty of our dataset,” states Schorn. Sipkema adds: “It was striking to realise that nearly all sponge species living at 150 m are different from the ones we see in the shallow water above.” COSMos analysed the metagenomes of five mesophotic sponges, their microbial symbionts, and their natural product biosynthetic potential. Thirteen more sponges have been sequenced since the project ended. From the original five metagenomes, 435 MAGs were assembled, representing 435 genomes from uncultured microbes. COSMos has thus given the scientific community an unprecedented look into the genomic capacities of mesophotic sponge symbionts. In addition to the genomic data, which will be made public, thousands of biosynthetic gene clusters have been identified.
From genomes to molecule production: heterologous expression
“For the first time, we have been able to look at the microbial communities of these understudied mesophotic sponges and discover new types of biosynthetic gene clusters. Connecting genes to their molecular products, we are beginning to decipher the chemical language of sponge inhabitants,” adds Schorn. Schorn is now focusing on the heterologous expression(opens in new window) of known and newly identified biosynthetic clusters. Heterologous expression from uncultured bacteria has rarely been done successfully. It would eliminate harvesting tremendous quantities of animals and enable sustainable large-scale production. “If we also discover a molecule with therapeutic applications, it will highlight the need to preserve understudied areas of the ocean like the mesophotic zone,” adds Schorn. Schorn and Sipkema are pushing the frontiers of marine natural products research and applications by revealing the genomic capacities hiding in enigmatic mesophotic sponges.