Simulating protein signalling
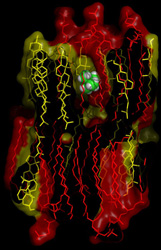
Signal transduction pathways are the mechanisms that allow living cells to respond to external stimuli such as hormones, neurotransmitters or growth factors. These molecules normally bind to membrane receptors and initiate a cascade of downstream intracellular events that lead to gene activation, cell proliferation or chemotaxis. Deregulation of signalling pathways is often the cause of many diseases including cancer and neurological disorders. Transduction of information through a single protein is known as allostery and is mediated by changes of protein shape and activity brought about by the binding of a molecular ligand. In allosteric proteins, it is clear that correlations between distant sites are encoded in the protein matrix. However, our knowledge on how this is achieved is currently limited. To this end, the ‘Protein signalling pathways elucidated via novel correlation analysis of molecular dynamics simulations’ (Protsign) project employed the technique of MD to study two important classes of signalling proteins: G-protein coupled receptors (GPCRs) and PDZ protein-binding domains. The MD technique was able to sample the structural flexibility of proteins and to simulate spontaneous allosteric protein transitions. Additionally, MD simulations were used to delineate solute permeation across biological membranes, a highly important process from biophysical, physiological and pharmacological perspectives. The same method proved useful for studying the chemistry of aerosols from seawater and how ions travelling with the water droplets play an important role in the chemistry of the atmosphere. Protsign demonstrated the applicability of the MD technique for simulating various scientific processes, especially in predicting the allosteric behaviour of certain proteins. This will help to dissect the molecular mechanisms involved during signal transduction and has broad scientific and medical significance.