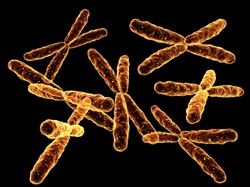
Chromosomes search for the perfect match
The 'In vivo choreography of DNA molecules and repair proteins during the search for homology' (CHOREOGRAPHY OF HR) project focused on one pathway activated when double-stranded breaks occur. Known as homologous recombination, the mechanism uses an undamaged identical DNA sequence to repair the broken chromosome. The researchers showed that when double-stranded breaks occur, the mobility of chromosomes increases dramatically. This increases the chances of finding a homologue for a matching sequence. To trace chromosome movement, the scientists used fluorescent dye to mark two homologous DNA regions in yeast cells that had a double set of chromosomes (diploid state). Before damage, the two homologous regions explored only 3 % of the space in the nucleus and occupied separate areas. Following double-strand break induction, however, homologous loci (stretches of DNA sequences) colocalise 10 times more often. As moving DNA can lead to unwanted chromosome arrangements, the team investigated the mechanisms involved. Using state-of-the-art high-resolution microscopy, they could image DNA movement up to 1 000 times faster. Project scientists found that DNA exploration is optimised. Fast, local motion over a short time scale gives precision and this is combined with large-scale motion on bigger time scales. Chromosome motion is diffusive, spreading out from a point gradually — this makes for an efficient search. CHOREOGRAPHY OF HR has already published a paper in Nature Cell Biology on speed of chromosome mobility and another is in preparation on the mechanisms involved. Data on this most basic of repair mechanisms could have huge significance for disease therapy generally.