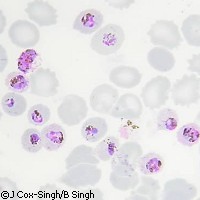
Scientists sequence genomes of two malaria parasites
Two international teams of scientists have sequenced the genomes of two major malaria parasites: Plasmodium vivax and Plasmodium knowlesi. Their results highlight the similarities and differences between the different Plasmodium species that cause malaria, and could lead to the development of new drugs and vaccines to tackle the deadly disease. Malaria is caused by parasites of the genus Plasmodium that are transmitted from human to human by certain mosquitoes. At least five species of Plasmodium are known to cause malaria in humans: P. falciparum, P. vivax, P. ovale, P. malariae and P. knowlesi. Some three quarters of cases and 90% of deaths from malaria are caused by P. falciparum whose genome was sequenced in 2002. In these latest studies, published in the journal Nature, scientists unravel the genomes of P. vivax, the second most deadly strain of the disease, and P. knowlesi, which is increasingly recognised as a major cause of malaria in humans, particularly in south-east Asia. P. vivax is the leading cause of malaria outside Africa, and some 2.6 billion people, mostly in Asia and Latin America, are at risk of contracting the disease. Unlike P. falciparum, P. vivax thrives in more temperate climates and during cooler seasons. Although it is rarely fatal, P. vivax infection is characterised by repeated episodes of illness which may go on for months. Widespread drug resistance means it is becoming increasingly difficult to treat cases of malaria caused by P. vivax. Analysis of the P. vivax genome reveals gene families that help the parasite break into red blood cells via previously unknown routes. Dormancy genes were also discovered; this information could be exploited to develop drugs designed to disrupt the parasite's dormant stage. 'Plasmodium vivax relapse presents serious challenges to scientists and doctors alike,' said Anthony Fauci of the National Institute of Allergy and Infectious Disease in the US. 'Completion of the P. vivax genome promises to provide new insights into the biology of vivax malaria and new leads of therapies and vaccines.' The second paper published in Nature outlines the genome of P. knowlesi. This strain's natural host is the kra monkey, but it is increasingly being found in humans, particularly in south-east Asia. Many human cases of malaria once attributed to P. malariae are now thought to be due to P. knowlesi; the two strains are difficult to tell apart using microscopy alone. As with P. vivax, the P. knowlesi genome brought to light a number of surprises. For example, it has genes which appear to mimic human genes involved in the regulation of the immune system. The scientists suspect that these parasitic genes help P. knowlesi interfere with the human host's ability to recognise infected blood cells. 'Our study demonstrates the power of sequencing additional malaria genomes to unravel as yet undiscovered and fascinating aspects of the biology of malaria parasites,' commented Dr Arnab Pain of the Wellcome Trust Sanger Institute, which is also working on sequencing the genomes of the remaining two species of Plasmodium that are known to infect humans. 'Unusually, the key genes that we think help the parasite to evade detection and destruction by host defences are scattered through the genome. In the other species we have examined, these genes are most often near the tips of the chromosomes.' 'This is our first view of a monkey malaria parasite genome. It brings us intrigues and surprises as well as new resources to help in the fight against malaria,' added Dr Alan Thomas of the Biomedical Primate Research Centre in Rijswijk, the Netherlands. 'P. knowlesi is closely related to the second-most common cause of human malaria, P. vivax. With our new understanding of the genetic architecture of both parasites, we will more efficiently translate our studies on P. knowlesi to other human parasites. Just as important, the genome will help in understanding human cases of knowlesi malaria.' EU support for the research on P. knowlesi came from the BioMalPar ('Biology and pathology of the malaria parasite') project, which is financed by the 'Life sciences, genomics and biotechnology for health' Thematic area of the Sixth Framework Programme (FP6) and VIRIMAL ('Structure, function and vaccine potential of the vir multigene family in malaria') project, which is funded through the 'Quality of life and management of living resources' Specific programme of the Fifth Framework Programme (FP5).