The gene prospectors searching for hidden life in hot springs
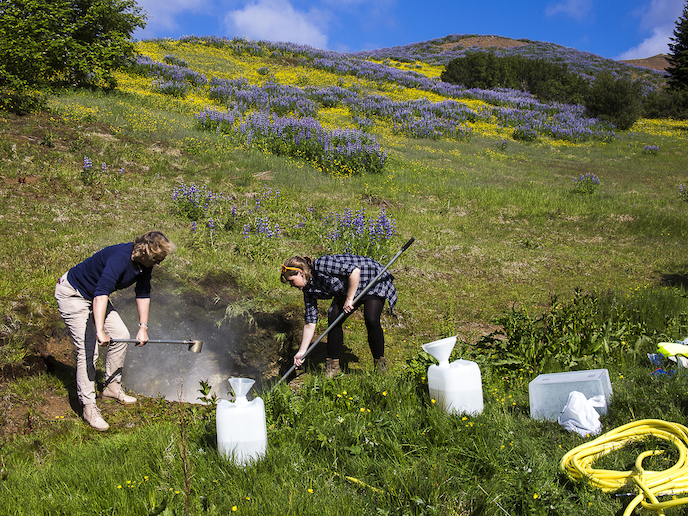
Gene products from viruses have often proven useful(opens in new window) for laboratory applications, specifically processing of DNA and RNA. Fourteen research institutes and commercial laboratories from eight different nations collaborated on the Virus-X(opens in new window) project to search for new genes to add to this toolkit. “Genes of viruses are still the largest unexploited sequence space to explore,” says Arnthor Ævarsson, Virus-X project coordinator. “There’s a huge reservoir of genes we haven’t seen before.” To find new genes, the researchers gathered water samples – up to tens of litres at a time – from ocean sites in Norway and hot springs in Georgia, Iceland and Japan. The locations were chosen because enzymes that can work at high temperatures are often suitable for use in lab-based applications. Samples were also taken from ocean sites in a search for enzymes active at lower temperatures. All the samples were taken through a series of filtration steps to concentrate the viral and bacterial gene fraction before this material was sequenced. “The fundamental concept in our approach is we do not use conventional methods of trying to grow viruses in the lab,” notes Ævarsson, a step that would require identifying and culturing the host bacteria. “Instead we isolate the genetic sequences from environmental samples in a metagenomic-based approach.” This is particularly important as a fraction of the genes discovered are associated with the candidate phyla radiation(opens in new window) – a large and mysterious group of bacteria that cannot be cultured in the lab, and are only known about through this metagenomic approach. Once the viral gene samples had been sequenced, powerful software was used to piece together these fragments into coherent lengths of DNA and RNA. Gene sequences of interest were identified and spliced into bacteria to synthesise the proteins they encoded. “We try to produce in sufficient quantities so we can determine three-dimensional structure using X-ray crystallography,” explains Ævarsson, a researcher at the independent Matis laboratory(opens in new window) in Iceland. “Once we have that, we have a much better chance of trying to figure out its function by structural comparison with known enzymes.” The results are still being investigated, although the team has already made several notable discoveries which are in the process of being published. But still they have merely scratched the surface, with only several hundred of some 50 million genes investigated. “We still have this treasure trove to work through,” says Ævarsson. The EUR 8 million endeavour – described by Ævarsson as his ‘dream’ project – was funded entirely through the EU’s Horizon 2020 programme. “This support was absolutely crucial, you can’t do a project of this scale and ambition without adequate funding,” he adds. “It’s a long pipeline, with a lot of people involved, with different kinds of expertise at all different levels.” Some of the gene products identified by Virus-X are already being developed for commercial use, and are likely to reach the market within the next few years. For Ævarsson, though, the thrill lies in the chase: “There is so much territory within the virome that has never been seen,” he concludes. “It’s like discovering new countries that nobody has been to before.”