Biophysical approach to tackling antibiotic resistance
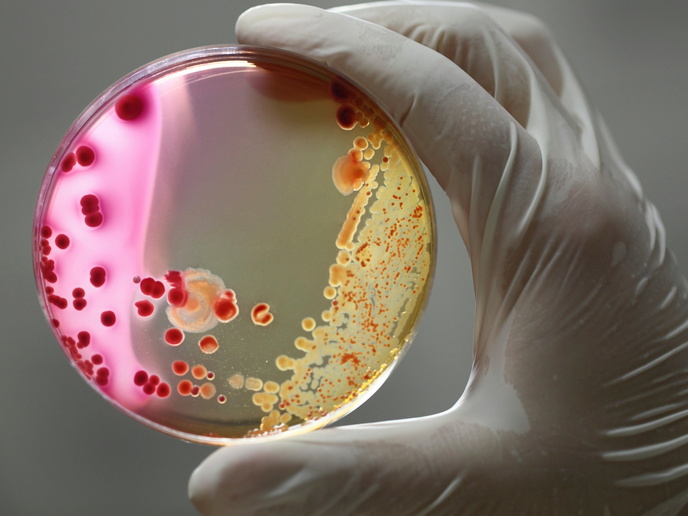
Since the discovery of penicillin in 1928, antibiotics(opens in new window) have been used to fight bacterial infections, both in hospitals and in the community at large. Every time a new antibiotic is discovered however, resistant bacterial strains emerge, rendering treatments ineffective. In this arms race between the medical and bacterial world, it would appear that humanity is on the losing side. “Many of the largest pharmaceutical companies, which were the driving force behind finding new antibiotics, have ended their antimicrobial programmes,” explains Tolerome project coordinator Nathalie Balaban(opens in new window), professor of Biophysics at the Hebrew University of Jerusalem(opens in new window) in Israel. “Uncovering new antibiotics is not seen as a good business model. Developing new drugs takes years. As a result, there are very few new antibiotics in the pipeline.” Some bacterial strains have become resistant to all antibiotics. Balaban warns of the dangers of the emergence and unchecked spread of an infectious and resistant strain.
Predicting bacterial evolution
The Tolerome project sought to address this looming health challenge through achieving a better understanding of how bacterial resistance evolves. This work began in the lab, through exposing bacteria to antibiotic treatments. “We were able to show that some evolving bacteria develop a property called tolerance,” says Balaban. “In this state, bacteria are dormant. We demonstrated that tolerance can be an important stepping stone towards antibiotic resistance, because tolerant bacteria are able to survive antibiotic treatment.” Balaban and her team also collaborated with Jerusalem hospitals to examine life-threatening bacterial blood infections. Bacterial strains were sequenced on a daily basis, enabling researchers to see bacterial evolution in action. “The same evolutionary trajectory we recorded in the lab was also occurring in the blood,” notes Balaban. “We were able to show that bacterial tolerance is a clinical problem and should be taken into account when treating bacterial infections.” A critical step was applying these findings to mathematical models, which then predicted how bacteria strains would evolve. “Putting your findings into equations can be a really powerful tool,” explains Balaban. “While it can sometimes be hard to see what is going on in the lab, mathematical modelling gives you a predictive understanding that is really powerful.”
More effective treatments
This pioneering combination of scientific evaluation and mathematical modelling holds the promise of enabling medical professionals to better predict the rapid evolution of antibiotic resistance. This in turn could lead to the prescription of more effective treatments. In particular, Balaban believes that the Tolerome project opens the door to the more efficient use of existing antibiotics. While finding and developing new compounds to treat bacterial infections can take years, there are a range of existing antibiotics that could be effective if used in combination. This would also give drugs already on the market a longer lifespan, by lowering their chances of becoming obsolete. “We now have this knowledge and understanding of bacterial evolution, as well as resistance and tolerance to antibiotics,” she says. “I am confident that being able to mathematically predict the trajectory of this evolution will help us in the future to identify combinations of drugs that can block this evolution, or at the very least, substantially delay the evolution of drug resistance.”