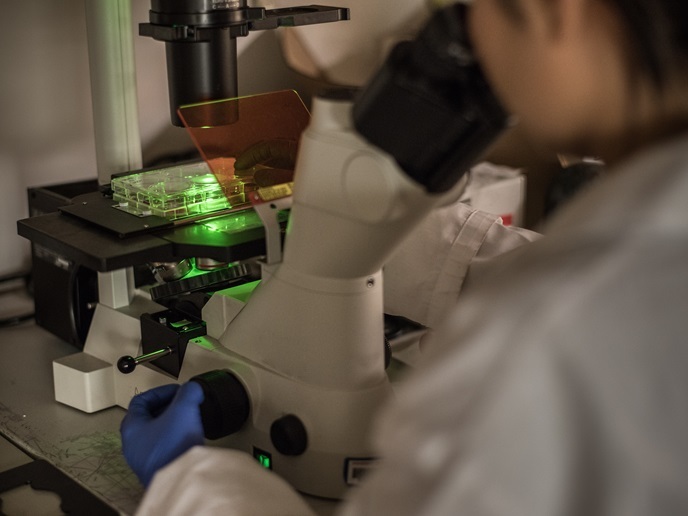
Label-free mapping of biomolecules in brain tissue
Imaging MS enables the label-free detection and mapping of a wide range of biologically relevant substances. It is a relative science, producing a normalised response with reference to a standard. Relative quantitation of pharmaceuticals is possible if the MS response factors (ratio analyte signal to the analyte quantity) are known. However, it is impossible to know a priori the MS response factor for every single peptide or protein. The EU-funded project ENIGMAS (Explicitly normalized imaging mass spectrometry) addressed this roadblock – the single biggest factor limiting clinical application of the powerful technique. Pioneering work led to development of a method for explicit normalisation of every peptide detected during imaging MS experiments. Team members utilised the recently released mouse model exploiting stable isotope labelling by amino acids in cell culture (SILAC). The SILAC technique for quantitative MS-based proteomics cultures cells in identical media with the exception of the amino acids present. One contains unlabelled amino acids and one has amino acids (usually lysine) in labelled form. The mouse version puts the labelled lysine in the animal's diet. Investigators used the technique to create a mouse reference standard in which all 12C6-lysine residues were replaced with 13C6-lysine. Tryptic digestion of the SILAC protein extract produced tryptic peptides containing a single 13C6-lysine. On-tissue digestion of normal mouse tissue produced 12C6-lysine-containing peptides. These were compared to the standard such that images of all tryptic peptides were normalised using their own isotopically labelled analogues. In addition to the sample preparation techniques, ENIGMAS developed a huge database of peptides and proteins from imaging MS of mouse brain tissue sections. An automated bioinformatics-based analysis pipeline took care of the rest. The technology was used to provide insight into the spatial and temporal biomolecular changes in mouse brain following an animal model of migraine. The new member of the imaging MS family of tools will have inestimable impact on increasing understanding in animal models of neurological diseases. Alignment of the imaging data sets with a genome-wide, high-resolution atlas of gene expression will further expand capabilities by linking biomolecular changes to gene expression.