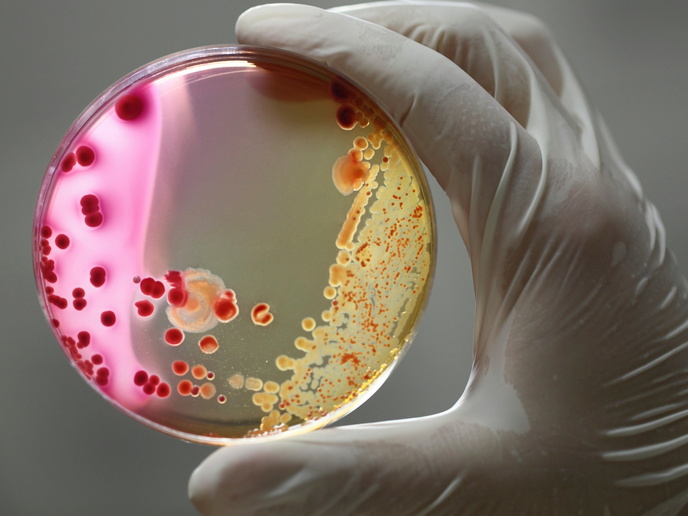
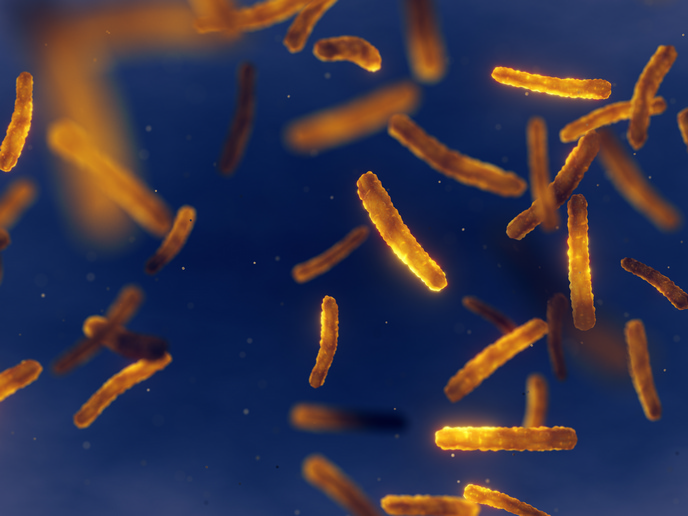
Bacteria that produce antibiotics
The quote, which Nobel-Prize winning physicist Richard Feynman famously scrawled on a chalkboard just before his death in 1988, read, "What I cannot create, I do not understand." Along these lines, EU-funded researchers of the PTM-FLEX (Synthetic biology approach for the design of new-to-nature peptide-based antibiotic molecules) project employed synthetic biology to design living systems that can produce desired bioactive molecules with increased flexibility. Their overall goal was to produce novel antibiotics such as lantipeptides from ribosomally synthesised and post-translationally modified peptides (RiPPs). Biologically active natural products, RiPPs can be produced by diverse organisms such as prokaryotes, eukaryotes and archaea. Through extensive post-translational modifications (PTMs) of ribosomally synthesised precursor peptides via selected enzymes, RiPPs with desired structure and functionality can be obtained. These RiPPs would have high stability, molecular target specificity and expanded functionality. Furthermore, the lower costs of genome sequencing have made it possible to rapidly and affordably identify selected RiPPs. PTM-FLEX researchers successfully established an E. coli expression platform to isolate and characterise promising RiPP producer strains. In addition, they developed a high-throughput screening method for the analysis of several potential antibiotic producers on a daily basis. Project activities have paved the way to discovery of new-to-nature antibiotics. The rapid clinical translation of such products will increase the availability of anti-bacterial drugs to treat or prevent bacterial infections.