Modelling anti-microbial resistance to help prevent malaria
Antimicrobial resistance (AMR) is one of the major looming challenges humanity faces today. A rise in the use of antibiotics over the past century, in both humans and animals, has led to a surge in pathogens resistant to antimicrobial drugs. AMR can develop if lower quality drugs are used, or if patients do not complete their course of treatment. The drug resistance of an infection is an evolutionary process, often requiring multiple mutations to occur, explains Tamsin Lee, a specialist in Resilient and Sustainable Systems for Health(opens in new window) at The Global Fund(opens in new window). “These resistant pathogens survive to be transmitted onwards, spreading resistance throughout the population,” she says. Each year, AMR causes thousands of deaths across the EU, and costs healthcare systems millions of euros. Yet forecasting its prevalence is challenging, in part because of the difficulties around data collection.
Modelling antimicrobial resistance
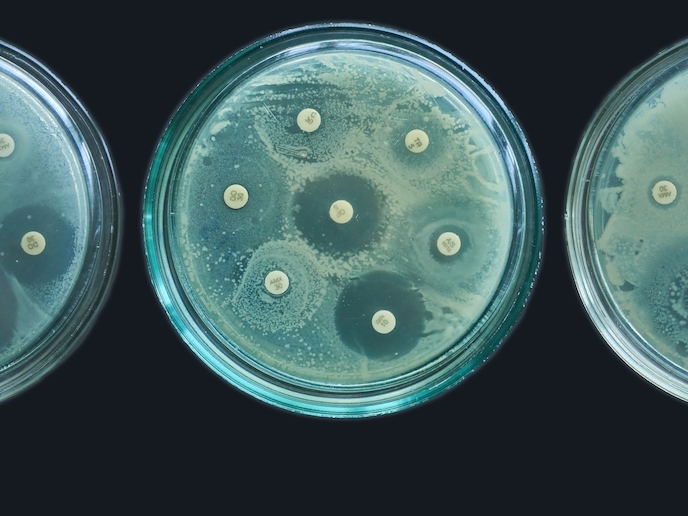
In the EU-funded EstAMR project, undertaken with the support of the Marie Skłodowska-Curie Actions programme(opens in new window), scientists worked to model the resistance of a parasite that causes malaria, Plasmodium falciparum. This has developed resistance to the latest antimalarials. The results could help forecast the frequency of drug-resistant malaria in different locations. The team considered socioeconomic status as an aggravating factor in the prevalence of drug resistance. “Perhaps the drugs are of a lower quality, or a patient shares their treatment with a family member,” Lee explains. To understand this link more clearly, researchers need lots of data. Drug resistant malaria infections are identified using molecular marker surveys, which identify mutations that are responsible for resistance. “Ideally, we could monitor where and when mutations occur in a region and track the spread of this variant throughout a population, enabling us to anticipate when and where it will next occur,” says Lee. But molecular marker surveys are expensive. This means only regions suspected of having drug resistant parasites are surveyed, meaning the data collected is biased.
Using a spatiotemporal model
To get around this problem, Lee and her team developed a spatiotemporal model, which allows for changes in space and time. They populated it with simulated data, considering a situation in the not-so-distant future, where drug resistant infections will be cheap and easy to test for. Using this simulated data, the EstAMR team was able to identify which health care centres were more likely to see greater ‘emergence’ of drug resistance. “These health care centres are probably using substandard drugs, and thus we can intervene and learn as to why they’re using substandard drugs,” explains Lee. “Perhaps the location is remote, making it difficult to keep drugs in stock, or perhaps their supplier is questionable, or perhaps there is a need to educate the patients, or make the drugs cheaper so that they finish the course,” says Lee.
Education to ensure full course of treatment is followed
The results of the work could be used to prevent the spread of drug resistant malaria and extend the lifespan of specific drugs. “Drug resistance is a real problem, but it’s unethical to withhold treatment, even if it’s substandard," says Lee. “To tackle drug resistance, we need to ensure that everyone has access to quality treatment and that they’re educated so as to finish the course of treatment,” she adds.