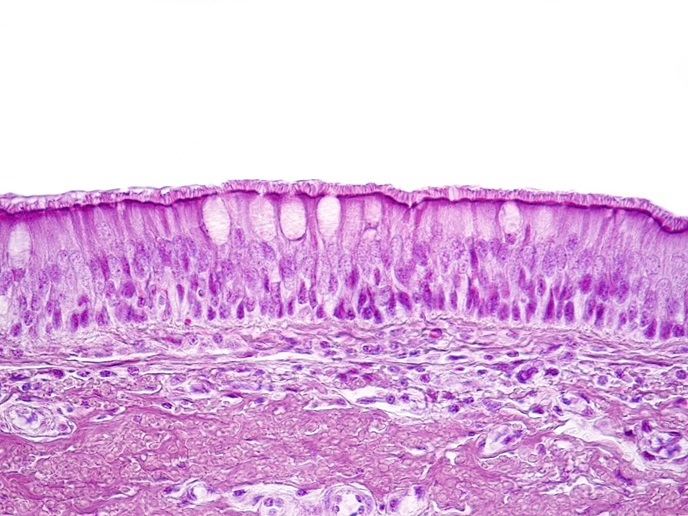
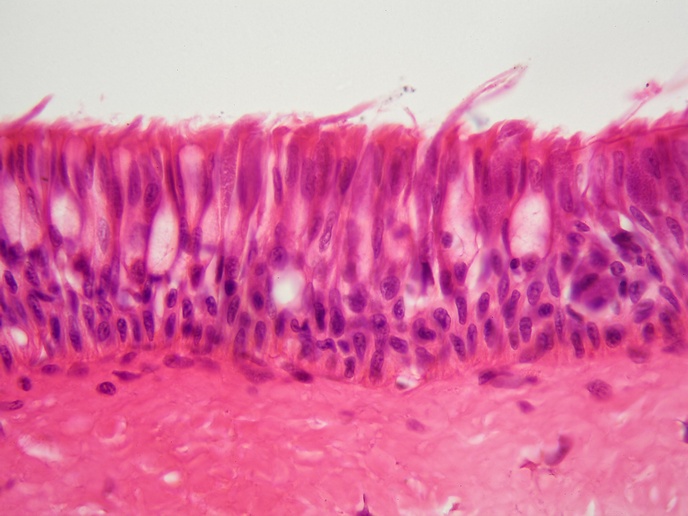
Cancer cell diagnostics with bacterial biosensors
Cancer remains one of the world’s leading causes of death, accounting for around 8.2 million fatalities each year. In many cases, mortality is driven not by the primary tumour but by metastasis, a process fuelled by circulating tumour cells (CTCs) that escape into the bloodstream. CTCs are phenotypically heterogeneous, and different subpopulations contribute in distinct ways to metastasis, therapy resistance and clinical outcome. Their detailed characterisation could transform tumour diagnostics and personalised treatment strategies. However, conventional methods struggle with low sensitivity, limited multiplexing capacity, and reliance on bulky laboratory infrastructure.
Converging different technologies into a single system
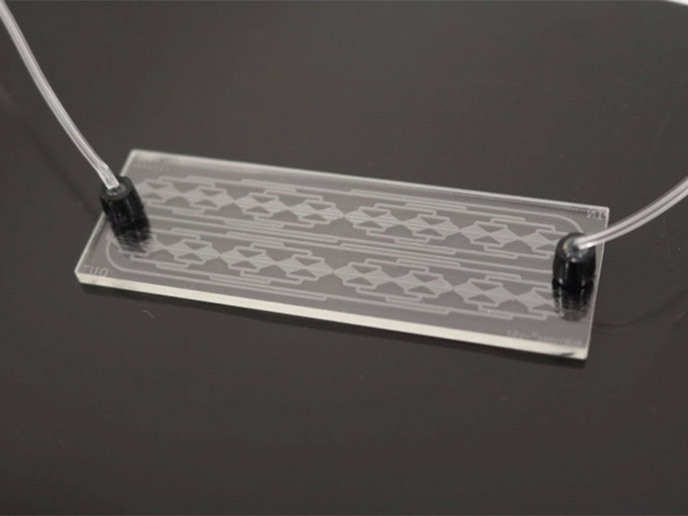
The EU-funded BIOCELLPHE(opens in new window) project set out to close this gap by developing a radically new approach to protein profiling of individual CTCs. “The vision is to transform how we detect specific proteins on the surface of individual tumour cells by integrating cutting-edge advances in nanotechnology, synthetic biology, microfluidics and artificial intelligence (AI) into a single, easy-to-use diagnostic platform,” explains project coordinator Isabel Pastoriza Santos. Cells express multiple proteins on their surface, which are often shared with other cell types. Therefore, the detection and identification of CTCs for cancer diagnosis requires measuring many protein biomarkers at once. To isolate CTCs, the consortium developed a lab-on-a-chip(opens in new window) device that passively sorts and traps cell-containing droplets from peripheral blood samples of cancer patients. These cells are subjected to downstream molecular analysis.
Engineered bacteria as living sensors
At the core of the concept are genetically engineered Escherichia coli strains designed to recognise specific protein biomarkers on the surface of CTCs. These strains carry receptors on their outer membrane, which enables selective attachment to validated cancer biomarkers. The specific adhesion of bacteria to target proteins on the cellular membrane triggers the production of Raman-active chemical compounds, which can be detected with ultrahigh sensitivity using surface-enhanced Raman scattering (SERS)(opens in new window), through their unique spectral fingerprints. To achieve this aim, the consortium engineered signalling and metabolic pathways to produce distinct Raman-active reporters, enabling simultaneous (i.e. multiplexed) detection of multiple proteins. Preliminary data from clinical analyses demonstrated that the engineered bacteria could detect and distinguish at least three different cell membrane proteins on CTCs. These findings highlight the potential of multiplex phenotyping in real patient samples.
Beyond the state of the art
While full integration into a single diagnostic device remains a future objective, BIOCELLPHE established a new scientific and technological framework for protein profiling at the single-cell level. The consortium’s multidisciplinary expertise in synthetic biology, nanotechnology, microfluidics, and AI enabled progress that had not previously been feasible. Future steps include optimisation, larger clinical validation studies and exploration of translational pathways. The BIOCELLPHE concept of employing engineered bacteria for sensing extracellular targets has the potential to advance precision oncology and biomedical diagnostics in general. “The foundations laid during the project open the door to applications well beyond cancer,” emphasises Pastoriza Santos.