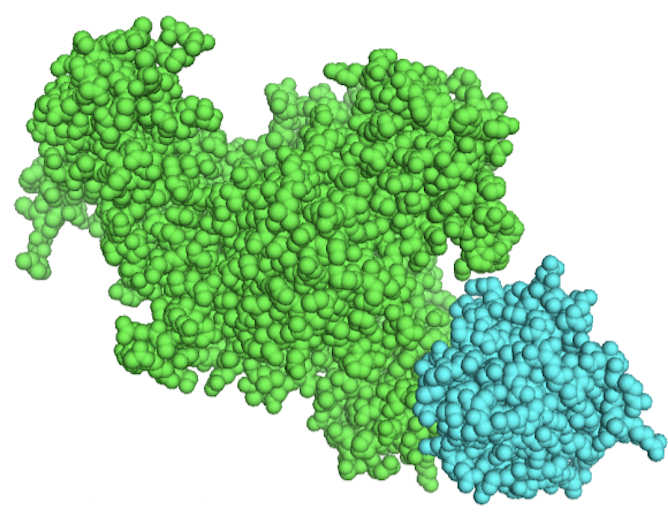
Shining a light on the relationship between protein sequence, structure and function
After water, proteins are the most abundant molecules in our bodies, making up muscles and other body tissues such as our hair. They are an essential component of all living organisms and scientists will be able to better understand them and develop medicines, thanks to new computational analysis lead by Marie Curie fellow Dr Lucy Colwell. Her team of mathematicians, chemists and biochemists at Cambridge University have discovered relationships between the sequences of proteins, their 3D structures and functions. “These findings will focus research in the future. The relationship between protein sequence and protein structure and function is one of the great problems of our time,” Dr Colwell, who worked on the Evo-Couplings project, says. New drug breakthroughs could be made more easily as a result of EVO-COUPLINGS as the team’s work improves scientists’ ability to predict the tertiary structure and the interaction partners(opens in new window) of a protein. The research, undertaken with the support of the Marie Curie programme, was a novel approach that was initially received skeptically by peers. But that has since changed. “I’m excited to see how mainstream these ideas have become,” she said. “Before our work it was much more difficult to predict protein structure and interactions from sequences alone. This approach is becoming standard in the field and has formed a key component of a number of recent important advances.” Cracking the matrix The researchers included mathematicians who developed methods that use random matrix theory – a probabilistic approach developed by physicists – to help the chemists and biochemists analyse protein sequence data. Experimental techniques in recent years have enabled natural scientists to gather large amounts of data for research but sifting through that information to find what is useful can be a headache. “My job is to cut through the noise,” said Dr Colwell. “To use protein sequences to predict structure, we had to first identify and remove the ‘noise’ in the data caused by the fact that different proteins are related to each other. This signal has to be ‘normalised’ out of the data before mathematical models can be built that make useful predictions.” Analysts use data visualisation methods to help identify structure in a range of fields: these methods can be applied to any type of data. Dr Colwell is now working with Google to use advances in machine learning to spot patterns that are difficult for humans to identify.