Can bacteriophages combat antibiotic resistance?
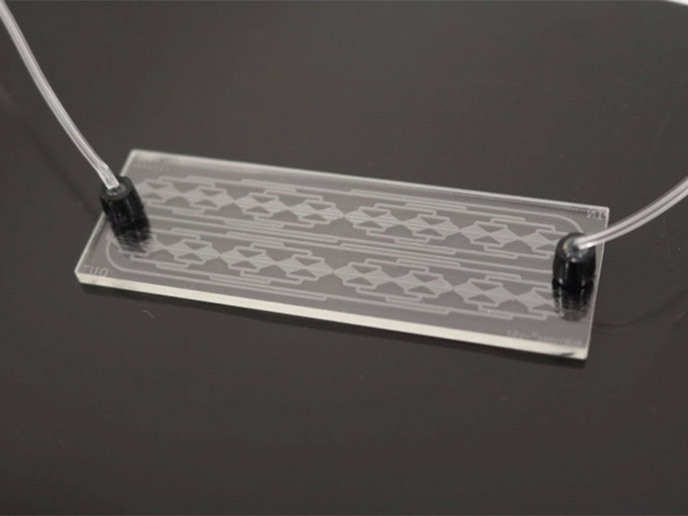
Bacteriophages are viruses that require a bacterial host to reproduce and they are capable of transferring genetic material between bacteria. Deciphering this interaction is central to developing effective strategies that involve phage and bacteriophage-based products. Investigating bacteriophage-bacteria interactions using microfluidics Supported by the Marie Curie programme, the EvoMachine-Phage(opens in new window) project was designed to understand the mechanisms underlying bacteria-bacteriophage antagonistic coevolution. The work focused on the phage-induced high cell density (PIHCD) phenotype previously observed by the Marie Curie fellow Dr Steven Quistad. As he explains, “the PIHCD phenotype was observed after mixing bacteriophages sampled from a local compost heap in Paris with the common soil bacterium Pseudomonas fluorescens.″ Although he expected the bacteriophage to either kill the bacteria or go into dormancy within the host chromosome, to his surprise the infected bacteria grew better. This observation urged the EvoMachine-Phage project fellow to investigate the underlying mechanism and identify the responsible bacteriophage. For this purpose, Dr Quistad exploited a novel microfluidic device created by MilliDrop(opens in new window), a spinoff biotech company founded at ESPCI Paris(opens in new window). The MilliDrop analyser is a fully-automated system for microorganism culture and analysis that allows the growth of bacteriophages and their hosts in thousands of small sub-microlitre droplets. The ‘evolution machine’ as Dr Quistad calls it, provides excellent bacterial growth conditions by keeping bacterial cells in constant motion and offers the capacity to select specific droplets of interest and analyse them. Using this device, Dr Quistad and colleagues investigated the impact of individual bacteriophages on host physiology and successfully identified multiple candidates responsible for the PIHCD phenotype. Ongoing work focuses on the identification of gene candidates that may be responsible for this phenotype. Future prospects of microfluidics in microbiology “Demonstrating the utility of microfluidics in the investigation of phage/host interactions was undoubtedly the most significant achievement of the EvoMachine-Phage project,″ outlines Dr Quistad. Additionally, the unparalleled opportunities for interdisciplinary collaboration offered by ESPCI Paris were central to the success of the project. The institute brought together chemists, physicists, engineers, and biologists while maintaining ties with industry. Employing microfluidics to rapidly test the impact of bacteriophages on host physiology could prove useful in a clinical setting. The gold standard plaque-based method requires approximately 48 hours for the isolation of bacteriophages from an environmental sample. However, when it comes to antibiotic-resistant infections, which can spread rapidly, expediting bacteriophage isolation and identification could significantly improve treatment outcome. The MilliDrop device has the potential to provide results of clinical significance faster. Furthermore, it could be employed as a tool in the battle against multi-drug resistance by testing the virulence of specific bacteriophage isolates against bacterial strains of interest. The EvoMachine-Phage project provided fundamental insight into bacteriophage physiology and their interaction with bacteria. Considering the involvement of bacteriophages in a range of biological processes from carbon cycling to the maintenance of human health, project findings are expected to find applications in various fields apart from bacteriophage-based therapeutics.