Evolution as it happens
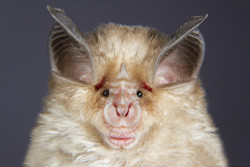
Researchers in the 'Dissecting speciation using a genomics approach' (RHINOSPEC) project have compiled a massive amount of data on one aspect of species interaction, inbreeding or hybridisation. Using new next genome sequencing (NGS) methods, they have identified genes that have introgressed, that is, they are expressed in the hybrid bat. The assumption is that genes important for survival will introgress and show in the phenotype. The study focused on co-distributed horseshoe bats in China, Rhinolophus pearsoni pearsoni (R. p. pearsoni), R. p. chinensis and R. yunanensis. Previous research has already indicated that introgression has occurred. The RHINOSPEC team determined to what extent this introgression had occurred. Data from two new NGS protocols, transcriptome sequencing (RNAseq) and targeted resequencing, yielded around 10 000 genes per group to characterise. Out of the 3 655 genes that were studied, 134 genes were either putative introgression or speciation genes, that is, involved in reproductive isolation. Identifying genes using a new RNA-based method has 'baited' 2 000 genes that have been resequenced. Eighty of these genes are involved in echolocation and the team will be able to test if this sensory method plays a part in reproductive isolation. A completely new study will focus on the importance of the X chromosome in speciation. Also, the significance of chromosome rearrangements in species mixing can be assessed. Two papers have already featured in peer-reviewed journals and another five are anticipated. RHINOSPEC has collected a large amount of data for analysis on exactly how species mix and become separated in the wild. In particular, knowledge will increase on the relevance of interbreeding in natural populations.