Novel players of mitochondrial gene regulation
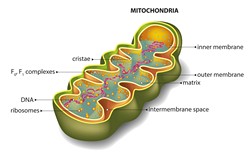
Mitochondria, the powerhouse organelles of the cell, possess their own genetic material, 80 % of which codes for mitochondrial RNA. However, several aspects of mitochondrial RNA biogenesis and regulation are incompletely understood. The FASTK (FAS-activated serine/threonine kinase) family of RNA-binding proteins have emerged as key regulators of RNA processing. In particular, loss of function mutations in FASTKD2 causes a rare form of mitochondrial encephalomyopathy. FASTKD2 seems to localise to and regulate mitochondrial homeostasis but the mechanisms remain elusive. Scientists on the EU-funded MOMEFAST (Molecular mechanisms employed by the newly assigned RNA-binding protein FASTKD2) project set out to investigate whether RNA binding of FASTKD2 contributes to the disease phenotype. To identify FASTKD2 targets, they aligned the FASTKD2-bound RNA target sites with the mitochondrial genome and discovered a set of mitochondrial transcripts including 16S ribosomal RNA (RNR2) and NADH dehydrogenase subunit 6 (ND6) mRNA. To validate results, scientists used RNA interference or gene editing to deplete FASTKD2 from cell lines. Extensive biochemical and metabolic analysis unravelled a pronounced global mitochondrial translation defect and an impairment in cellular respiration. In particular, aberrant processing and expression was seen for the mitochondrial RNR2, concomitant with strong expression of 7S RNA, a small non-coding RNA associated with perturbed homeostasis. Taken together, the MOMEFAST work identified previously unknown aspects of the molecular network of the RNA-binding protein FASTKD2. These will help comprehend the mechanism underlying mitochondrial encephalomyopathy and maybe other diseases involving RNA-binding proteins.