Immune system IDs pathogenic fungi
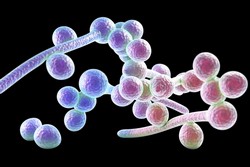
Ca is a dimorphic fungus that grows as yeast and filamentous cells; one of the few species of the Candida genus that causes the infection candidiasis in humans. Innate immune cells tolerate commensal colonisation of Ca, yet deal effectively with opportunistic mucosal and systemic invasion of pathogenic Ca. The innate immune cells and fungi interact predominantly at the level of the fungal cell wall, recognising pathogen-associated molecular patterns (PAMPs) by pattern recognition receptors (PRRs). The cell wall composition of virulent Ca hyphae (clusters of branch-like fungus cells) differs from Ca yeasts resulting in differential immune recognition and signalling. The goal of the EU-funded FUNGISORT (Sorting out host recognition of fungal infection) project was to study PRR interactions with fungal PAMPs and to develop new tools for pathogenic state recognition. Sortagging (sortase-mediated transpeptidation) is a chemoenzymatic system for site-specific labelling of proteins with small probes that permit the attachment of affinity handles (like biotin or fluorophores). The technique can be used to label purified proteins or to perform immunoprecipitation experiments with proteins expressed on the surface of living cells. FUNGISORT researchers applied sortagging for the study of the C-type lectin receptor DC-SIGN (dendritic cell-specific intercellular adhesion molecule-3-grabbing non-integrin) that plays an important role in innate immune recognition of Ca. Using sortagging, the carbohydrate recognition domain of DC-SIGN was fluorescent-tagged and used in pull down and confocal microscopy experiments in the macrophages. Researchers identified glycoproteins that are secreted by Ca and interact with DC-SIGN. These results indicate the need to investigate if these glycoproteins could be found in vivo and whether they have DC-SIGN-mediated immunomodulatory functions. The results of the FUNGISORT study contributed significantly to the understanding of immune recognition of fungal infection. They also presented new avenues for the development of diagnostic tools to determine fungal pathogenicity.