Cell migration data analysis and storage revamped
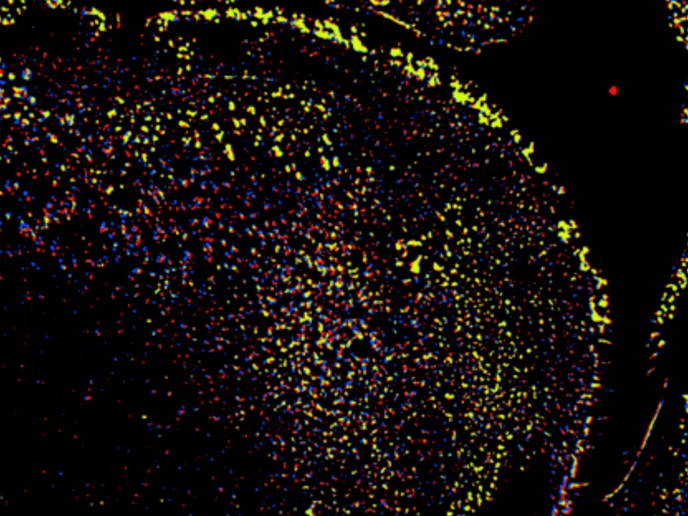
Cell migration is a fundamental process involved in development and morphogenesis, immune function, wound healing and cancer metastasis. Over the last few years, advances in cell migration research have enabled widespread use of multiplexing and multi-parameter post-processing of cell migration analyses. As it is one of those areas that generates Big Data for analysis and storage, a largely unmet bioinformatics need has emerged. “The MULTIMOT project has risen to this enormous challenge and developed both standards and a repository,” summarises Dr Lennart Martens, leader of the Computational Omics and Systems Biology group in Flanders Institute for Biotechnology, Belgium, the coordinating institute. Report, share and reposit The community-driven standards creation was guided by the Cell Migration Standardisation Organisation (CMSO)(opens in new window). This has resulted in minimal reporting requirements for a cell migration experiment as well as controlled vocabularies and the biotracks format for cell migration data. “All development can be found (and joined!) on the CMSO GitHub pages(opens in new window),” notes Dr Martens. The repository for cell migration data is also available online(opens in new window). Significantly, the first large-scale data sets derived from MULTIMOT are being made available now (pending acceptance of the corresponding manuscripts). “Moreover, MULTIMOT has also devised novel ways to look at multi-scale cell migration data, and these new analysis workflows have already been applied in two research initiatives for personalised medicine,” outlines Dr Martens. One involving cancer treatment is based on patient-derived tumour cells, and the other is providing patient-specific diagnoses based on peripheral blood leukocyte motility. Challenges turn out to be opportunities “Bringing such an interdisciplinary group of experts together to create standards, software, standard operating procedures and novel analysis software was a significant challenge on the face of it,” comments Dr Martens. Throughout the project, this turned out to be an easy and quick exchange of information, experience and data. The high calibre of scientists also meant that issues with experimental hiccups and protocol standardisation difficulties were thankfully anticipated by the experienced scientists and addressed swiftly without interrupting the flow of information or the overall progress of the project. “As the saying goes,” says Dr Martens, “a good team really is half the win!” Next steps in cell migration data analysis “The work of establishing a widely adopted open data exchange ecosystem in the field has only just kicked off in earnest with these developments,” Dr Martens emphasises. The research leader is of the opinion that with these crucial foundations now firmly in place, the future for open data sharing and thus enhanced value extraction from cell migration data looks very bright indeed. While the data sharing is freely available, this does not preclude the development of novel commercial endeavours centred on bringing the benefits of an open data exchange ecosystem to end users. “This has been an explicit part of MULTIMOT from its inception, with its SME inclusion, to ensure the relevance of its output for commercial companies, and for new commercial opportunities,” Dr Martens points out. In any life science, there is therefore an enormous opportunity for novel knowledge to be uncovered through reanalysis and repurposing of the collected data. With the MULTIMOT project, the field of cell migration has now joined other fields such as genomics, proteomics and metabolomics in establishing an open data exchange ecosystem to realise the full potential of the data.