Protein shape affecting mutation dictates activation in some childhood leukaemias
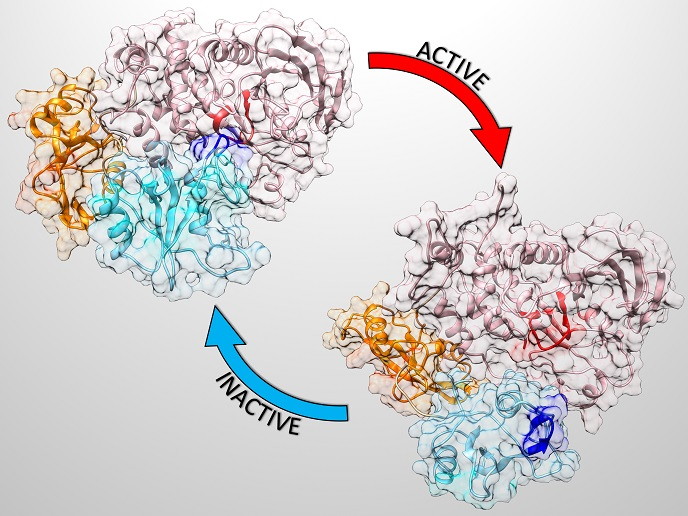
Proteins are complex molecules that play critical roles in the body. The SHP2 protein upregulates cellular processes, so much so that its mutations cause severe developmental disorders like the Noonan syndrome(opens in new window). Mutations in the SHP2 gene also give rise to juvenile myelomonocytic leukaemia and are more rarely found in other haematologic malignancies and tumours. SHP2 mainly exists in an autoinhibited (inactive) form, and it is only fully activated upon specific association with its binding partners. See image above of the very different forms of inactive and active protein.
Protein function and drugs to control mutations
“The MARS project has uncovered, at atomic level, the circumstances governing activation of SHP2. In addition, we built an atomistic model that explains how certain disease-associated mutations perturb such a mechanism,” outlines project coordinator, Prof. Jochen Hub. Achieving these objectives is an important prerequisite for the design of drugs that interfere specifically with SHP2 mutants and modulate their function. N-SH2 is the protein domain or area that recognises the all-important binding partners and blocks the SHP2’s active site in the autoinhibited structure. “We found that N-SH2 adopts two distinct conformations or shapes, only one of which is activating,” explains Dr Massimiliano Anselmi, the project’s Marie Skłodowska-Curie fellow. Interestingly, the researchers also found that the activation is not merely promoted by the simple binding of a ligand(opens in new window), as previously believed, but instead by shifting the population of N-SH2 from the non-activating to the activating conformation or shape. This means the ligand sequence and even its particular way of arranging into the binding site are crucial aspects in selecting the activating conformation of N-SH2.
Digestive problems and challenges
As the simulations increasingly supported the model of N-SH2 activation, the team realised that most of the previously available experimental data could not be unequivocally interpreted. “Despite the fact nothing actually contradicted our conclusions, nothing seemed to definitely confirm them,” Dr Anselmi recalls. The key problem, he explains, “is that N-SH2–activating peptide sequences are also partly digested (more precisely: de-phosphorylated) by N-SH2, which makes a quantitative interpretation of the experiments very challenging.” The answer came from a mutation reported more than 10 years ago. “We found that the structural effects of this pathogenic mutation affect the region of N-SH2 that rearranges during the activation and could be clearly explained uniquely using our model,” he enthuses. “It was an astounding and decisive piece of work,” adds Prof. Hub.
Clarifying the fuzzy and onward steps to understanding the activation mechanism
Knowledge of the activation mechanism of SHP2, and the possibility of inducing it artificially in lab experiments enables further questions to be answered. Dr Anselmi cites just one example: “Using small-angle X-ray scattering (SAXS)-derived data, it will be possible to identify the three-dimensional structure of the open state of SHP2, further validating the activation pathway proposed.” Conclusions provided by the MARS project have challenged and rejected several commonly held views on SHP2 activation. “Often the huge amounts of data reported in literature are scattered and somehow fuzzy, and a model helps to organise and settle them in a rational way,” summarises Prof. Hub.