The breakdown of clumped tau proteins to cure Alzheimer’s disease
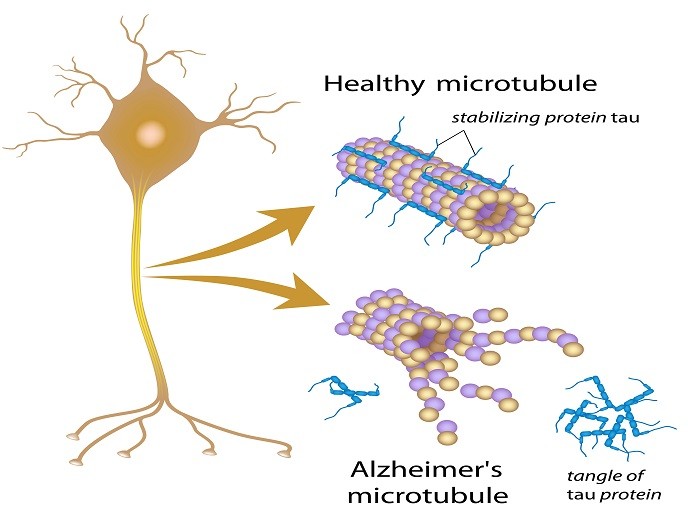
According to the World Health Organization, more than 47 million people worldwide are affected by dementia, with Alzheimer’s disease being the most common and widespread among them (60-70 %). One feature that Alzheimer’s patients have in common is the presence of rod-like structures known as amyloid fibrils. Composed of a protein tau in its aggregated form, these have also been implicated in a number of other neurodegenerative diseases (tauopathies). These include frontotemporal dementia, progressive supranuclear palsy and primary age-related tauopathy. In healthy neurons, the tau protein stabilises microtubules that help guide nutrients from the cell neuron body to the axon and dendrites. In Alzheimer’s disease, however, abnormal chemical changes cause tau to detach from microtubules and stick to other tau molecules, forming threads that eventually join to form tangles inside neurons. These tangles block the neuron’s transport system, which harms the synaptic communication between neurons. Despite major research efforts, the mechanism of tau aggregation and its pathogenic consequences are not yet understood. Therapeutics to cure these tauopathies, including Alzheimer’s disease, are urgently needed.
Untangling tau
Saurabh Gautam, post-doctoral Marie Skłodowska-Curie research fellow with the REVERSING TAUOPATHY project outlined the objective of the initiative: “Our goal at the Max Planck Institute of Biochemistry(opens in new window) was to find out the conditions for dissolution of such protein aggregates in the human cellular model of tau ‘clumping’ or aggregation.” The mechanism and cellular machinery involved in the dissolution of these tau aggregates could lead to the development of lead compounds for new drugs. Preliminary work identified two main conditions when tau aggregates were disassembled. Starvation of the cell model using a medium devoid of amino acids – Earle's Balanced Salt Solution – caused aggregate breakdown. Switching off the expression of the tau protein also caused disaggregation of the rogue protein. The researchers then used state-of-art techniques such as stable isotope labelling with amino acids in cell culture – SILAC(opens in new window) based mass spectrometry, to identify the components of the cellular machinery involved in the dissolution of tau aggregates. “After screening various conditions with this popular method for quantitative proteomics, we were able to show that intracellular tau aggregates can be disassembled in the cells, a huge breakthrough,” Gautam declares.
Motivation overcomes the technical challenges
“This was a challenging project from the start since there have been no reports of dissolving the intracellular tau aggregates to our knowledge,” explains Gautam. “However, the potential use of findings and outcome for potential therapeutic applications motivated us to work in this direction,” she emphasises. Another major roadblock was replicating the findings from the intracellular environment inside a test tube. This had to be accomplished by purifying various identified cellular components using recombinant DNA technology and trying to dissolve tau aggregates in vitro. “This will allow us to pinpoint the exact mechanisms and reactions going on,” Gautam explains. However, the work is still in progress. “We at REVERSING TAUOPATHY believe that answers will contribute to finding a cure for these tauopathies,” he concludes.