Glycans could be the key to immunity-boosting therapies
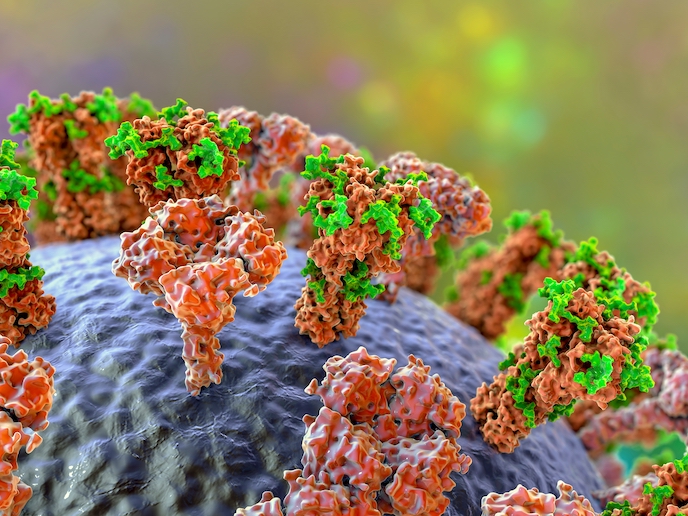
Glycans(opens in new window) are carbohydrate-based polymers ubiquitous within living cells. Key to many biological processes, they enable cells to recognise molecules and orchestrate cell-to-cell interaction. Within cells, they are involved in protein folding, positioning, transport and quality control. Viruses and bacteria not only attach themselves to glycans, but the spike protein of viruses, including SARS-CoV-2, is sometimes coated with glycans, camouflaging the virus from the immune system. Glycan changes often correlate with pathological states, such as inflammation and cancer. The structure of glycans is not genetically encoded, but rather results from the actions of various glycan-processing enzymes, making their position and composition hard to determine. Additionally, their complexity makes manipulation difficult using current methods. The SWEETOOLS project, funded by the European Research Council(opens in new window), developed a methodology to construct libraries of glycans linked to a peptide. These ‘glycopeptides conjugates’ can be used to screen various glycan-processing enzymes. “Knowing how specific enzymes bind with and process glycans, means our library’s glycopeptides can be tweaked to control the functioning of specific glycan-processing enzymes, paving the way for new drug therapies,” explains project coordinator Milan Vrabel.
The glycopeptide library
Synthetic peptide libraries are short chains of amino acids – 4 to 15 in SWEETOOLS – prepared using a chemical technique known as the one-bead-one-compound(opens in new window) methodology. A set of polymer resin beads containing an amino acid are divided into equal portions to which the next amino acid is added. These portions are pooled and chemically coupled to the next amino acid. The beads are divided again and the steps repeated, producing a synthetic library where each bead contains a unique sequence of amino acids – a peptide. SWEETOOLS added a small molecular inhibitor that binds to selective enzymes, resulting in peptide libraries of different amino acids but the same inhibitor. “While the peptides recognise, select and bind to the enzyme, the inhibitor stops that enzyme functioning. We can then monitor any resulting changes, such as whether inhibition of this enzyme boosts the immune system,” adds Vrabel. The peptide library was built using click reactions – a set of fast, efficient and selective chemical reactions. Glycan residue was then added to the peptides. The resulting glycopeptide library can be used to find proteins, such as lectins, that recognise specific glycans. As this is the mechanism by which bacteria and viruses bind to their targets, understanding it offers the prospect of subverting it. For cellular experiments, probes were developed which produced light as a result of key chemical reactions. This enabled the team to track and visualise the biomolecules in detail. “Preliminary data shows that one of our glycopeptides protects T-cells from Galectin 1(opens in new window) triggered cell death, a mechanism certain cancer cells use to evade our immune system. With the benefit of this glycoconjugate, we could fight these cancers more effectively,” says Vrabel.
Towards a future of ‘living drugs’
The team also discovered that synthetic versions of naturally occurring glycans can be metabolically processed by live cells, forming natural glycoconjugates. The team engineered these to be ‘chemically’ modifiable inside cells or at the surface, offering cheaper immunomodulators. “Reagents could install artificial molecules onto living cells to protect them, with cellular interactions prevented or new interactions promoted. We could install antibodies onto immune cells which then find and destroy cancers or protect against side effects after organ transplantation,” concludes Vrabel. The first patent application on the chemical re-engineering of cells has already been filed, with in vitro experiments using different cancer cell lines underway.